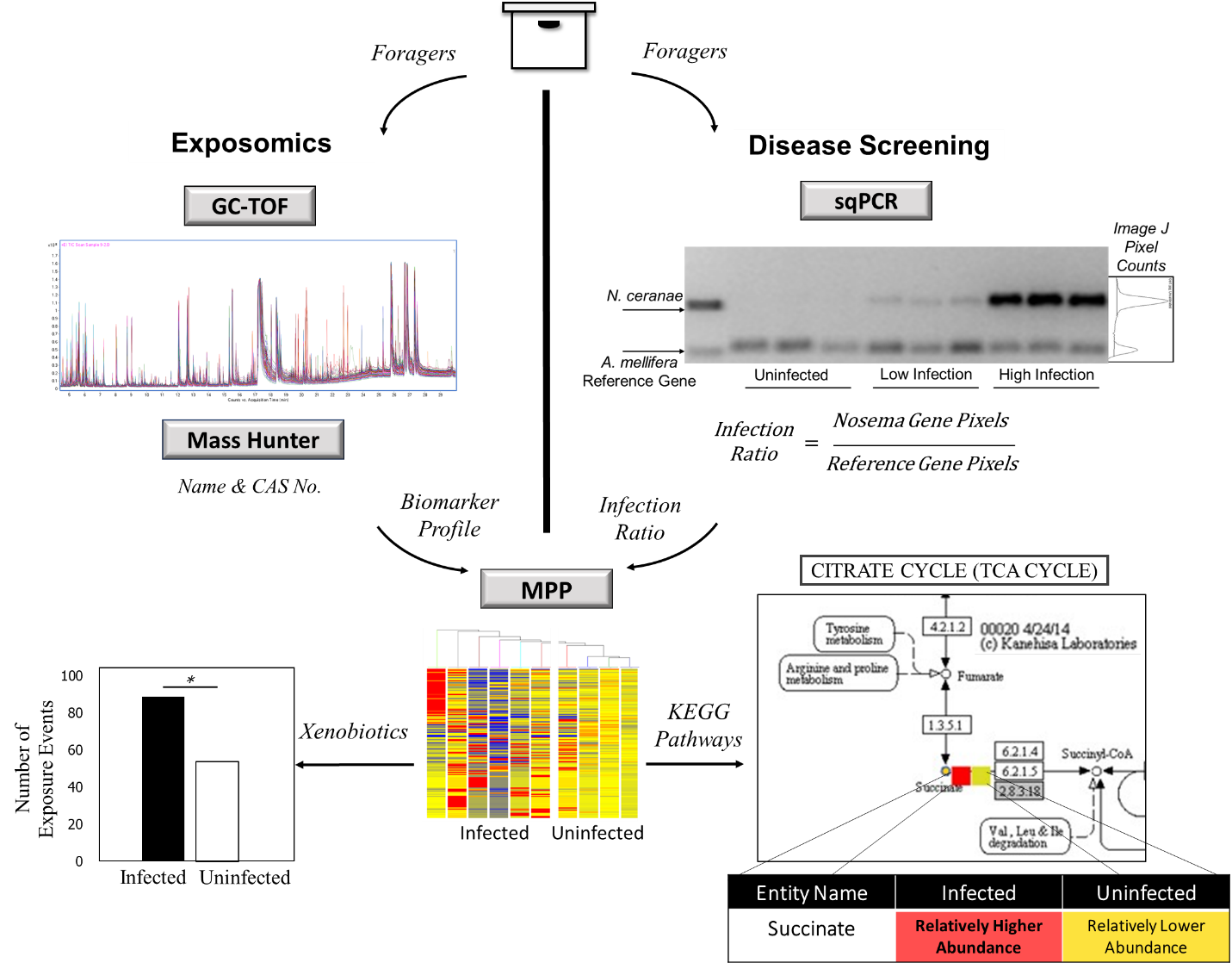
Honey bees are an integral part of the global agricultural economy as they pollinate roughly one third of the food that we eat everyday. Within the past decade, globally bee health has been in decline, and a multitude of stressors including parasites, pesticide exposure, and losses of foraging habitat have been associated with this observation. Exposomics characterizes the plethora of chemicals in and around an organism and attempts to associate these with biological response pathways and chronic diseases. Using the systems biology framework, the exposome paradigm will be integrated with other 'omic' platforms to look for associations between health and unhealthy bee hives. Using metabolomics, transcriptomics, and bioinformatic pathway analysis, we will correlate exposomes with disease exposure identified by molecular techniques from bee extracts and link this to chemicals in the ambient hive air to identify novel robust chemical biomarkers for prognosis and/or diagnosis of hive health. This knowledge will be used to develop a cost effective, non-invasive technology for beekeepers to monitor hive health and identify risk of colony collapse.
Students will gain experience in wet lab molecular techniques such as semi-quantitative PCR, qPCR, and generating new molecular protocols for disease screening. Students will gain exposure to statistical data design and analysis. Bioinformatics for metagenomic, metabolomic, and KEGG pathway analyses will be used. These are skill sets and current approaches that are being used in the biomedical field for solving complex disorders such as cancer and Alzheimer's.
About Project Supervisors
Dr. Christopher Mayack
Assistant Professor
Molecular Biology, Genetics and Bioengineering Program
Faculty of Engineering and Natural Sciences
Sabanci University
Orhanli, Tuzla, Istanbul 34956 Turkey
Office: FENS 2061
Phone: +90-216-568-7038
E-mail: cmayack@sabanciuniv.edu
Fax: +90-216-483-9550
.jpg)